PCNN Fold 4
Back to PCNN overview · Back to main index
Fold metrics
| Metric | Value |
|---|---|
| Test accuracy | 11.46% |
| Test F1 score | 0.0872 |
| Hierarchical loss | 0.79275930 |
| P-adic loss (total) | 1506.07650000 |
| P-adic loss (mean) | 0.55309457 |
| Prime base | 79 |
| Hidden layer size | 27 |
| Max tags | 32 |
| Training samples | 10,836 |
| Test samples | 2,723 |
P-adic loss breakdown
| Agreement | Count | Share | Cost per mistake | Total contribution |
|---|---|---|---|---|
| Exact match | 312 | 11.46% | 0.000000 | 0.000000 |
| p^3 | 117 | 4.30% | 0.000002 | 0.000237 |
| p^2 | 318 | 11.68% | 0.000160 | 0.050953 |
| p^1 | 476 | 17.48% | 0.012658 | 6.025316 |
| p^0 | 1,500 | 55.09% | 1.000000 | 1500.000000 |
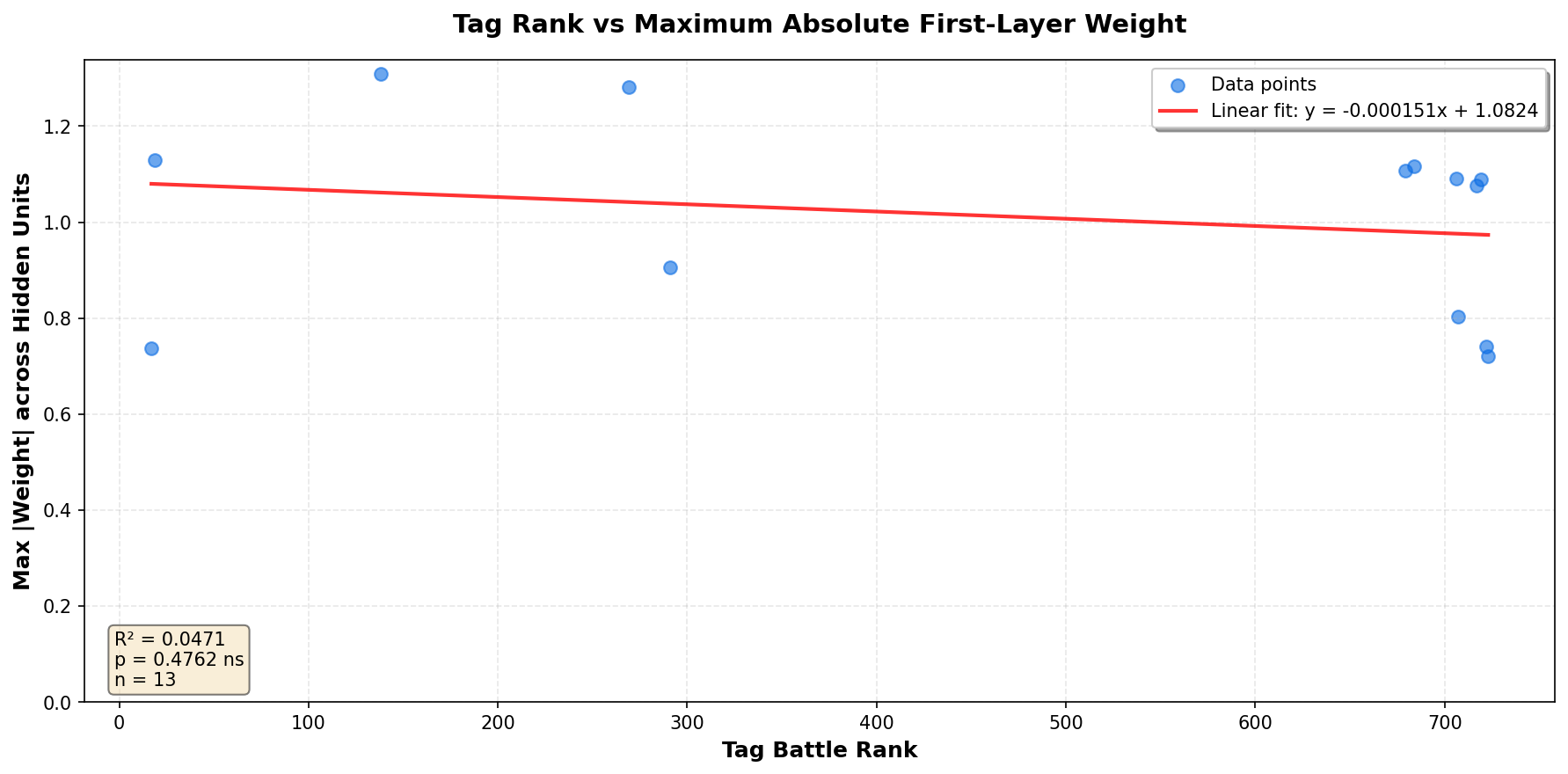
Tag Rank vs First-Layer Weight Magnitude
About p-adic loss
P-adic loss measures the distance between predicted and true taxonomy using p-adic metric (base 79). Lower values indicate closer predictions in the taxonomy hierarchy. This metric is shared with the umllr model for comparison.